High Dimension Data
High Dimensional Data
$$
\begin{bmatrix}
\mathbf{x}_ 1 \\
\mathbf{x}_ 2 \\
\vdots \\
\mathbf{x}_ N
\end{bmatrix} =
\begin{bmatrix}
x_1^{(1)} & x_1^{(2)} & \cdots & x_1^{(d)} \\
x_2^{(1)} & x_2^{(2)} & \cdots & x_2^{(d)} \\
\vdots & \vdots & \ddots & \vdots \\
x_N^{(1)} & x_N^{(2)} & \cdots & x_N^{(d)}
\end{bmatrix}
$$
Mean and Covariance of High-Dimensional Data
When working with high-dimensional data, it is important to understand the mean and covariance matrix, which are essential statistical measures that summarize the data’s location and spread.
1. Mean Vector
For a dataset with $ n $ samples and $ d $ features (dimensions):
- Let $ \mathbf{X} \in \mathbb{R}^{n \times d} $ be the dataset, where each row $ \mathbf{x}_i \in \mathbb{R}^d $ represents a data point and each column corresponds to a feature.
Definition:
The mean vector $ \mathbf{\mu} \in \mathbb{R}^d $ is defined as:
$$
\mathbf{\mu} = \frac{1}{n} \sum_{i=1}^n \mathbf{x}_i
$$
Element-Wise:
The $ j $-th element of the mean vector is:
$$
\mu_j = \frac{1}{n} \sum_{i=1}^n x_{ij}, \quad j = 1, 2, \dots, d
$$
Where $ x_{ij} $ is the $ j $-th feature of the $ i $-th sample.
2. Covariance Matrix
The covariance matrix $ \mathbf{\Sigma} \in \mathbb{R}^{d \times d} $ captures the pairwise relationships between features.
Definition:
$$
\mathbf{\Sigma} = \frac{1}{n-1} \sum_{i=1}^n (\mathbf{x}_i - \mathbf{\mu})(\mathbf{x}_i - \mathbf{\mu})^T
$$
Element-Wise:
The $ (j, k) $-th entry of the covariance matrix is:
$$
\Sigma_{jk} = \frac{1}{n-1} \sum_{i=1}^n (x_{ij} - \mu_j)(x_{ik} - \mu_k)
$$
Where:
- $ \Sigma_{jk} $: Covariance between feature $ j $ and feature $ k $.
- $ x_{ij} $: Value of the $ j $-th feature for the $ i $-th sample.
Properties:
- $ \Sigma_{jj} $: Variance of feature $ j $.
- $ \Sigma_{jk} $: Correlation between features $ j $ and $ k $ if scaled by their standard deviations.
- The matrix $ \mathbf{\Sigma} $ is symmetric: $ \Sigma_{jk} = \Sigma_{kj} $.
3. Matrix Representation
Using matrix notation, the mean vector $ \mathbf{\mu} $ and covariance matrix $ \mathbf{\Sigma} $ can be computed efficiently:
Mean Vector:
$$
\mathbf{\mu} = \frac{1}{n} \mathbf{X}^T \mathbf{1}
$$
Where:
- $ \mathbf{X}^T $: Transpose of the data matrix.
- $ \mathbf{1} $: A column vector of ones with size $ n $.
Covariance Matrix:
$$
\mathbf{\Sigma} = \frac{1}{n-1} (\mathbf{X} - \mathbf{1} \mathbf{\mu}^T )^T (\mathbf{X} - \mathbf{1} \mathbf{\mu}^T)
$$
Summary of Notation
| Symbol | Meaning |
|---|---|
| $ \mathbf{\mu} $ | Mean vector of the dataset ($ d $-dimensional). |
| $ \mathbf{\Sigma} $ | Covariance matrix ($ d \times d $). |
| $ \mu_j $ | Mean of the $ j $-th feature. |
| $ \Sigma_{jk} $ | Covariance between feature $ j $ and $ k $. |
Example of the Covariance
The Iris dataset has been successfully loaded. Here’s a brief look at the dataset:
| Sepal Length (cm) | Sepal Width (cm) | Petal Length (cm) | Petal Width (cm) |
|---|---|---|---|
| 5.1 | 3.5 | 1.4 | 0.2 |
| 4.9 | 3.0 | 1.4 | 0.2 |
| 4.7 | 3.2 | 1.3 | 0.2 |
| 4.6 | 3.1 | 1.5 | 0.2 |
| 5.0 | 3.6 | 1.4 | 0.2 |
Let’s compute the mean and covariance matrix for this dataset and visualize their insights.
Results from the Iris Dataset:
Mean Vector:
The mean of each feature (measured in centimeters) is:
- Sepal Length: $ 5.843 $
- Sepal Width: $ 3.057 $
- Petal Length: $ 3.758 $
- Petal Width: $ 1.199 $
Covariance Matrix:
The covariance matrix is:
$$
\mathbf{\Sigma} =
\begin{bmatrix}
0.6857 & -0.0424 & 1.2743 & 0.5163 \\
-0.0424 & 0.1900 & -0.3297 & -0.1216 \\
1.2743 & -0.3297 & 3.1163 & 1.2956 \\
0.5163 & -0.1216 & 1.2956 & 0.5810
\end{bmatrix}
$$
Interpretation:
-
Diagonal Entries:
- These are the variances of the features:
- Variance of Sepal Length: $ 0.6857 $
- Variance of Sepal Width: $ 0.1900 $
- Variance of Petal Length: $ 3.1163 $
- Variance of Petal Width: $ 0.5810 $
- These are the variances of the features:
-
Off-Diagonal Entries:
- These represent covariances between pairs of features:
- Positive covariance (e.g., $ 1.2743 $ between Sepal Length and Petal Length) suggests a positive relationship.
- Negative covariance (e.g., $ -0.3297 $ between Sepal Width and Petal Length) suggests a negative relationship.
- These represent covariances between pairs of features:
The Covariance Matrix with Species Encoding is as follows:
| Feature | Sepal Length (cm) | Sepal Width (cm) | Petal Length (cm) | Petal Width (cm) | Species Encoded |
|---|---|---|---|---|---|
| Sepal Length (cm) | 0.6857 | -0.0424 | 1.2743 | 0.5163 | 0.5309 |
| Sepal Width (cm) | -0.0424 | 0.1900 | -0.3297 | -0.1216 | -0.1523 |
| Petal Length (cm) | 1.2743 | -0.3297 | 3.1163 | 1.2956 | 1.3725 |
| Petal Width (cm) | 0.5163 | -0.1216 | 1.2956 | 0.5810 | 0.5973 |
| Species Encoded | 0.5309 | -0.1523 | 1.3725 | 0.5973 | 0.6711 |
Key Observations:
-
Species Encoded Relationships:
- Positive covariance with features like petal length ($1.3725$) and petal width ($0.5973$).
- Indicates that these features strongly vary with the species.
-
Feature Variability:
- Variance (diagonal values) is high for petal length ($3.1163$), meaning it varies most across the dataset.
- Sepal width ($0.1900$) has the least variance.

What can we get from this results?
As you can see, Petal Length has the largest variance (var = 3.1163). Meanwhile, it also has the highest covariance with species (1.37). This indicates that much of its variance is explained by the species, making Petal Length a potentially good feature for species classification.
Transformations
High-dimensional data transformation refers to the process of modifying or converting data that exists in a high-dimensional space (i.e., data with a large number of features or variables) into a more manageable or meaningful representation. This transformation can involve reducing dimensions, re-organizing data, or mapping it to a different space while preserving important information or relationships.
-
Source Dataset (${x}$) and Target Dataset (${m}$):
- The target dataset ${m_i}$ is generated by applying a rotation and translation to the source dataset:
$$
m_i = A x_i + b
$$ - Here, $A$ is the rotation matrix, and $b$ is the translation vector.
- The target dataset ${m_i}$ is generated by applying a rotation and translation to the source dataset:
-
Mean Transformation:
- The mean of the transformed dataset (${m}$) can be expressed as:
$$
\text{mean}({m}) = A \cdot \text{mean}({x}) + b
$$
- The mean of the transformed dataset (${m}$) can be expressed as:
-
Covariance Transformation:
- The covariance matrix of the transformed dataset (${m}$) is derived as:
$$
\text{Covmat}({m}) = A \cdot \text{Covmat}({x}) \cdot A^\top
$$ - This shows how the covariance matrix of the source dataset transforms under a linear transformation.
- The covariance matrix of ${x}$ is defined as:
$$
\text{Covmat}({x}) = \frac{1}{N} \sum_i (x_i - \text{mean}({x}))(x_i - \text{mean}({x}))^\top
$$
- The covariance matrix of the transformed dataset (${m}$) is derived as:
Eigenvector and Eigenvalue
Imagine you’re analyzing data (like in machine learning or physics). Eigenvalues and eigenvectors can:
- Find patterns: In large data (like PCA), eigenvectors show the “directions” of most variation, and eigenvalues tell how important each direction is.
- Simplify problems: Diagonalization makes hard matrix computations easier.
1. Eigenvector ($u$) and Eigenvalue ($\lambda$)
- An eigenvector $u$ of a matrix $S$ is a vector that does not change direction when $S$ is applied to it. Instead, it is scaled by a factor $\lambda$, the eigenvalue:
$$
S u = \lambda u
$$- $S$: A square matrix.
- $u$: An eigenvector (non-zero vector).
- $\lambda$: The corresponding eigenvalue.
2. Symmetric Matrices ($S$)
- If $S$ is symmetric ($S = S^\top$), it has special properties:
- All eigenvalues are real.
- Eigenvectors corresponding to distinct eigenvalues are orthogonal:
$$
u_i \perp u_j \quad \text{if} \quad i \neq j
$$ - Eigenvectors can also be normalized to form an orthonormal set ($|u| = 1$).
3. Orthonormal Matrix ($U$)
- By stacking all the eigenvectors of $S$ as columns into a matrix $U$:
$$
U = [u_1, u_2, \dots, u_d]
$$- $U$ is an orthonormal matrix, meaning:
$$
U^\top U = I \quad \text{(identity matrix)}
$$
- $U$ is an orthonormal matrix, meaning:
4. Eigenvalues as a Diagonal Matrix ($\Lambda$)
- Arrange eigenvalues $\lambda_1, \lambda_2, \dots, \lambda_d$ into a diagonal matrix:
$$
\Lambda =
\begin{bmatrix}
\lambda_1 & 0 & \dots & 0 \\
0 & \lambda_2 & \dots & 0 \\
\vdots & \vdots & \ddots & \vdots \\
0 & 0 & \dots & \lambda_d
\end{bmatrix}
$$
5. Diagonalization
- If $S$ is symmetric, it can be diagonalized using its eigenvectors and eigenvalues:
$$
S = U \Lambda U^\top
$$- $U$: Matrix of eigenvectors.
- $\Lambda$: Diagonal matrix of eigenvalues.
6. Key Properties of Diagonalization
- Simplifies computations, e.g., powers of $S$:
$$
S^k = U \Lambda^k U^\top
$$- $\Lambda^k$ is simply the diagonal matrix with each eigenvalue raised to the power $k$.
- Used in many fields such as:
- Principal Component Analysis (PCA).
- Solving differential equations.
- Modal analysis in engineering.
Principal Component Analysis (PCA)
Principal Component Analysis (PCA) is a statistical technique used to reduce the dimensionality of data while retaining as much variance as possible. It identifies new axes, called principal components, which are uncorrelated and align with the directions of maximum variance. PCA transforms the data to these new axes, ranks the components by their variance (eigenvalues), and allows dimensionality reduction by selecting the top components.
PCA in 3 Steps (More Accurate Breakdown):
-
Transformation (Centering the Data):
- Before applying PCA, you need to center the data by subtracting the mean of each feature. This step ensures that the principal components (axes of maximum variance) pass through the origin.
- Mathematically:
$$
x_{\text{centered}} = x - \text{mean}(x)
$$
-
Rotation (Find Eigenvalues and Eigenvectors):
- The goal of PCA is to find the directions (principal components) where the data has the most variance.
- This involves computing the eigenvectors and eigenvalues of the covariance matrix:
$$
S = \frac{1}{n} X^\top X
$$- The eigenvectors represent the new axes (principal components).
- The eigenvalues indicate how much variance is captured by each axis.
- Rotation refers to aligning the data along the directions of these principal components.
-
Dimensional Reduction (Keep Principal Components):
- After identifying the principal components, you can choose the top $k$ components with the highest eigenvalues (the directions of the most variance) and ignore the rest.
- This step reduces the dimensionality while retaining as much information as possible.
1. Original Dataset:
- The dataset ${x}$:
- It has $d$ features (dimensions).
- Each data point is a vector in a $d$-dimensional space.
2. Step 1: Covariance Matrix:
- The covariance matrix captures how features are correlated. It is computed as:
$$
\text{Covmat}({x}) = \frac{1}{N} \sum_{i=1}^N \left( x_i - \text{mean}({x}) \right) \left( x_i - \text{mean}({x}) \right)^\top
$$ - PCA works by diagonalizing this covariance matrix.
3. Step 2: Eigen Decomposition:
- Decompose the covariance matrix into eigenvalues ($\lambda$) and eigenvectors ($u$):
$$
U^\top \text{Covmat}({x}) U = \Lambda
$$- $U$: Matrix of eigenvectors (principal components).
- $\Lambda$: Diagonal matrix of eigenvalues (variance explained by each principal component).
4. Step 3: Choose $s$ Principal Components:
- Eigenvalues represent the variance explained by each principal component. They are sorted in descending order.
- To reduce dimensions:
- Choose the top $s$ eigenvalues that explain the most variance.
- Often, the ratio is calculated:
$$
\frac{\sum_{j=s+1}^d \lambda_j}{\sum_{j=1}^d \lambda_j}
$$- This ratio helps decide $s$ by ensuring the remaining variance (error) is small.
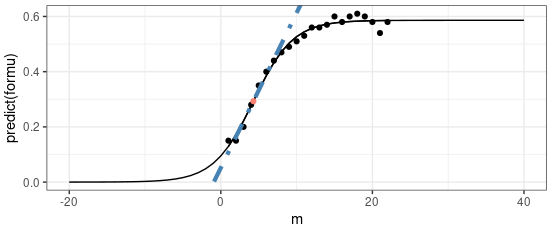
- Plotting $\lambda_i$ vs. $i$ (as shown in the slide) can help visualize where most variance is captured (the “elbow” point).
5. Step 4: Project Data to Lower Dimensions:
- Once you have selected $s$ principal components, project the original data onto this lower-dimensional space:
$$
\hat{x}_ i = \sum_ {j=1}^s \left[ u_ j^\top (x_ i - \text{mean}({x})) \right] u_j + \text{mean}({x})
$$- Here:
- $u_j$: The eigenvectors corresponding to the top $s$ eigenvalues.
- $\hat{x}_i$: The low-dimensional representation of $x_i$.
- Here:
6. Visualization from the Slide:
- The graph shows the eigenvalues ($\lambda_i$) vs. their indices ($i$):
- The blue curve represents the sorted eigenvalues.
- The orange circle highlights the “elbow” point, which suggests the optimal number of principal components to retain.
Summary of PCA Calculation:
- Center the data (subtract the mean).
- Compute the covariance matrix.
- Find eigenvalues and eigenvectors of the covariance matrix.
- Choose the top $s$ eigenvalues to decide the number of principal components.
- Project the data onto the top $s$ eigenvectors to get a reduced representation.
High Dimension Data